bioinformatictools.blogspot.com
bioinformatictools.blogspot.com
Bioinformatics Tools: In-silico characterization of proteins
http://bioinformatictools.blogspot.com/2011/11/in-silico-characterization-of-proteins.html
Bioinformatics has evolved as a great tool for molecular biologists. There are various tools available for helping in reducing the time required to analyze biological materials be it DNA, RNA, Proteins etc. I wish to list here few of the commonly used tools. Please send me suggestions to improve the content. If you like or dislike something, let me know, your inputs matters. contact me: drsanjivk[at]gmail[dot]com. Saturday, November 5, 2011. In-silico characterization of proteins. Different combinations ...
 russelllab.org
russelllab.org
Services offered by Protein Evolution
http://www.russelllab.org/services.shtml
This tool lets you interrogate data from our collaborative project on the Ciliary proteome, where we identified nearly 5000 interactions and over 50 complexes, involving more than 1000 proteins in the cilia. From Boldt, van Reeuwijk, Lu, Koutroumpas et al Nat Commun. From Betts et al, Nucl Acids Res. A database of protein-chemical interactions predicted by structure superimpositions: if you are interested in chemicals that might bind a particular protein or vice versa, then check it out. From Stark et al...
 ub.cbm.uam.es
ub.cbm.uam.es
Unidad de Bioinformatica CBMSO
https://ub.cbm.uam.es/home/links.php
Structural stability of ecological systems. Proteomics research: http:/ www.proteomicsresearch.org. Phyre: http:/ www.sbg.bio.ic.ac.uk/ phyre/. Fugue: http:/ www-cryst.bioc.cam.ac.uk/ fugue/. Robetta: http:/ robetta.bakerlab.org/. BlastProfiler: http:/ manaslu.aecom.yu.edu/blastprofiler.html. Hamap: http:/ trantor.bioc.columbia.edu/hmap/index.html. Mammoth: http:/ ub.cbm.uam.es/servers/mammoth/mammoth.php. Mammoth-mult: http:/ ub.cbm.uam.es/servers/mammoth/mammothmult.php.
 disprot.org
disprot.org
Disprot - Database of Protein Disorder
http://www.disprot.org/predictors.php
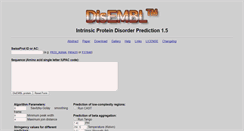
Xue, B., R. L. DunBrack, R.W. Williams, A.K. Dunker, and V. N. Uversky (2010) "PONDR-Fit: A meta-predictor of intrinsically disordered amino acids." Biochim. Biophys. Acta 1804(4):996-1010. Internal pointer to the PONDR-FIT meta-predictor and other PONDR methods. Linding R, Jensen LJ, Diella F, Bork P, Gibson TJ, Russell RB. "Protein disorder prediction: implications for structural proteomics." Structure. 2003;11(11):1453-9. Dosztanyi Z, Csizmok V, Tompa P, Simon I. "IUPred: web server for the predic...
 tango.crg.es
tango.crg.es
Tango
http://tango.crg.es/links.jsp
A computer algorithm for prediction of aggregating regions in unfolded polypeptide chains. Links to related sites. An algorithm to predict the helical content of peptides. Fragmenting the protein space. Program for calculating the folding pathways of proteins and the effect of a point mutation on the protein stability. A Cell Network Simulation Program. Predicting amylogenic regions in protein sequences. Prediction of protein-protein interAction of moDular domAiNs. Molecular Phenotypes of Human Diseases.
 smart.embl.de
smart.embl.de
SMART: Main page
http://smart.embl.de/index2.cgi
Schultz et al. (1998). Proc Natl. Acad. Sci. USA. Letunic et al. (2014). You may use either a Uniprot. Sequence identifier (ID) / accession number (ACC) or the protein sequence itself to perform the SMART analysis service. Sequence ID or ACC. Examples: Q4SNE7 Q4SNE7 TETNG nMLRLSLLLLLHCHWASSTLLGWVESPGYPHGYLPHASLNWSRCARKGHIISIRLVHLDLEKSRNCENDAVKVLSDGSPISILCGKKSFAELQSSVNPLLRSSTGGCLTLLFASDYSNTRRHSGFRGFYTEQDFDECRDDPEASCNQFCHNFVGGYYCSCRHGYHLKEDQRTCTVRCSEDLSGLREGALSSPSWPAPYPAHAHCLYTLSVDEHLQVELHFSDGFDVEQSTDGHCVD...
 landscape.syscilia.org
landscape.syscilia.org
Services offered by Protein Evolution
http://www.landscape.syscilia.org/services.shtml
This tool lets you interrogate data from our collaborative project on the Ciliary proteome, where we identified nearly 5000 interactions and over 50 complexes, involving more than 1000 proteins in the cilia. From Boldt, van Reeuwijk, Lu, Koutroumpas et al Nat Commun. From Betts et al, Nucl Acids Res. A database of protein-chemical interactions predicted by structure superimpositions: if you are interested in chemicals that might bind a particular protein or vice versa, then check it out. From Stark et al...
 mechismo.russelllab.org
mechismo.russelllab.org
Services offered by Protein Evolution
http://www.mechismo.russelllab.org/services.shtml
This tool lets you interrogate data from our collaborative project on the Ciliary proteome, where we identified nearly 5000 interactions and over 50 complexes, involving more than 1000 proteins in the cilia. From Boldt, van Reeuwijk, Lu, Koutroumpas et al Nat Commun. From Betts et al, Nucl Acids Res. A database of protein-chemical interactions predicted by structure superimpositions: if you are interested in chemicals that might bind a particular protein or vice versa, then check it out. From Stark et al...
 lindinglab.science
lindinglab.science
Digital ‘Rosetta Stone’ Decrypts How Mutations Rewire Cancer Cells — Rune Linding : Research Group
http://www.lindinglab.science/news
Only in current section. Digital ‘Rosetta Stone’ Decrypts How Mutations Rewire Cancer Cells. Scientists have discovered how genetic cancer mutations systematically attack the networks controlling human cells, knowledge critical for the future development of personalized precision cancer treatments. Lori Waters, Waters Biomedical, 2015. Computational Rosetta Stone decodes Mutations Rewiring Cancer Signaling. Lead researcher on the projects, Prof Dr Rune Linding from the Biotech Research and Innovation Cen...
 lindinglab.org
lindinglab.org
Tony Pawson (1952 - 2013) — Rune Linding : Research Group
http://www.lindinglab.org/news/tony-pawson-1952-2013
Only in current section. Tony Pawson (1952 - 2013). Tony Pawson (1952 - 2013). The entire world-wide research community mourns the passing of Dr Tony Pawson. It is with great sadness that we learned that we have lost our great hero, close friend, fantastic mentor, superb colleague, wonderful human being and maverick of a scientist Tony Pawson. Across biology and life-sciences this will be felt deep and long. There is no field we can imagine that will not be touched by this massive loss. Take care and sta...